Pocket Descriptors
Comprehensive analysis of protein pocket properties and characteristics
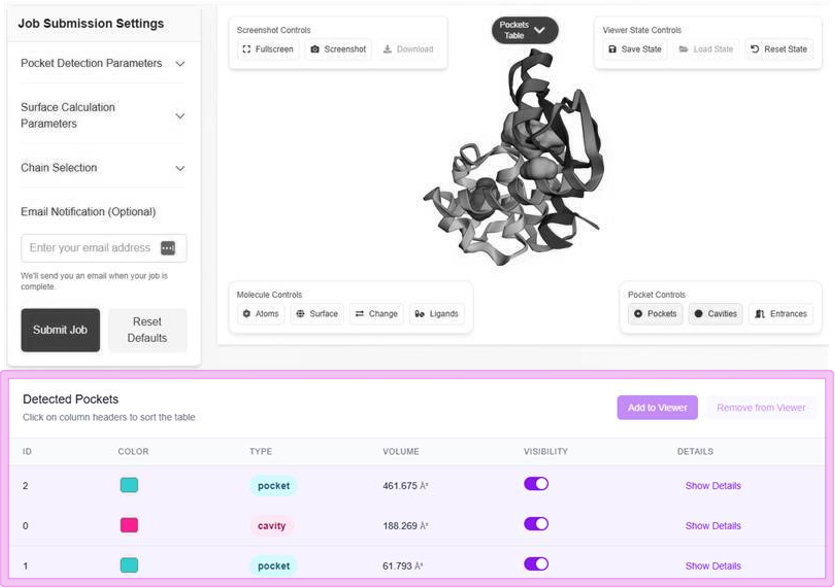
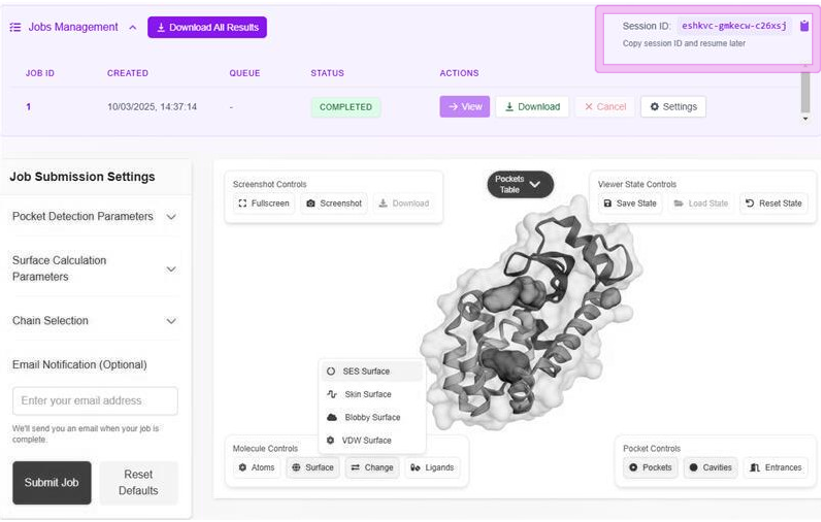
After NanoShaper delivers the set of pockets of a protein, a further script is used to estimate the molecular DrugPred descriptors with minor modifications (the entrance area computed by NanoShaper is used as an additional descriptor).
Pocket Descriptors
Basic Geometric Properties
Fundamental measurements of pocket geometry and shape
Surface Properties
Hydrogen bonding and hydrophobic characteristics
Hydrogen Bond Donor Areas
Hydrogen Bond Acceptor Areas
Hydrophobic Properties
Amino Acid Composition
Distribution and occurrence of different amino acid types
Detailed Description
Surface Properties
Each pocket identified by NanoShaper is characterized by descriptors from Krasowski et al. (2011), along with the entrance area provided by NanoShaper. Binding site properties include hydrogen-bond donor and acceptor surface areas (dsa_t and asa_t), calculated around polar atoms.
Area Calculations
The hydrophobic surface area (hsa_t) is derived by subtracting the hydrogen-bond donor and acceptor areas from the total surface area. Relative descriptors (dsa_r, asa_r, and hsa_r) are computed by normalizing each by the total binding site surface area, with the relative polar surface area (psa_r) defined as the sum of dsa_r and asa_r. Finally, the compactness descriptor is used to characterize cavity shapes.
Amino Acid Properties
The remaining descriptors, related to amino acid composition, were calculated based on the occurrence of different amino acid classes grouped by their physicochemical properties. Moreover, the occurrence of each amino acid of type (in_X) is reported as descriptors, computed as the sum of all the surface areas surrounding the amino acid X.