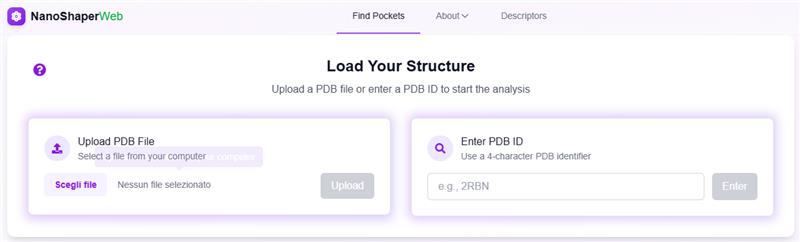
NanoShaperWeb
A powerful tool for analyzing molecular surfaces and predicting functional sites in protein structures
In structural biology and biophysical modeling, analyzing molecular surfaces and predicting functional sites in protein structures are crucial tasks. Evaluating molecular properties such as volume, surface area, and protein cavities is essential for understanding protein function and contributing to structural drug design.
Core Functionality
NanoShaper is designed to measure molecular properties, construct a Finite Divergence grid for solving PDE, and compute protein pockets. The algorithm works by triangulating the molecular surface using ray-casting to inspect the surface's inner regions in a grid-based space. This process generates an in/out map, allowing for cavity identification, volume estimation, and optional cavity filling.
Volume Estimation
The volume estimation is achieved through a triple ray-casting process, where rays are cast from the x, y, and z axes of the cube containing the molecule. The three resulting volume measurements are averaged for a highly accurate estimate.
Surface Calculation
For molecular surface area calculation, NanoShaper implements a parallel Marching Cubes algorithm. This algorithm triangulates the surface and ensures precise surface area measurements. When voids are removed, the triangulation reflects this by producing a consistent surface representation without cavities.
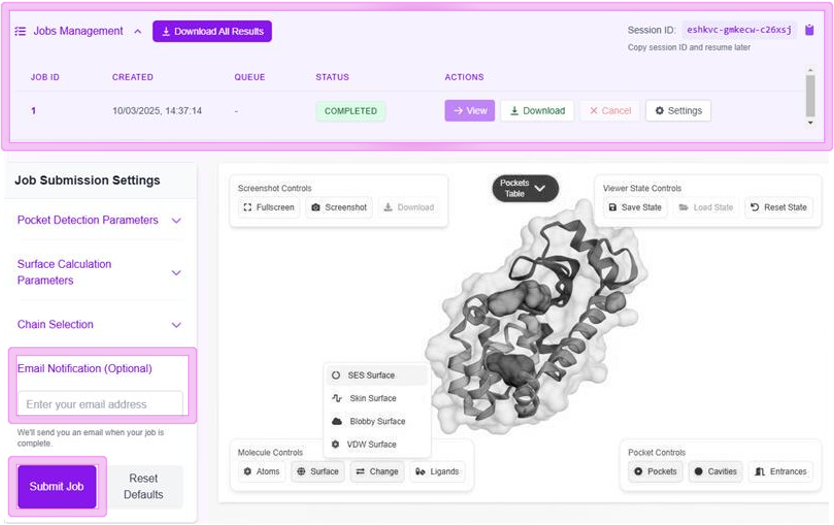
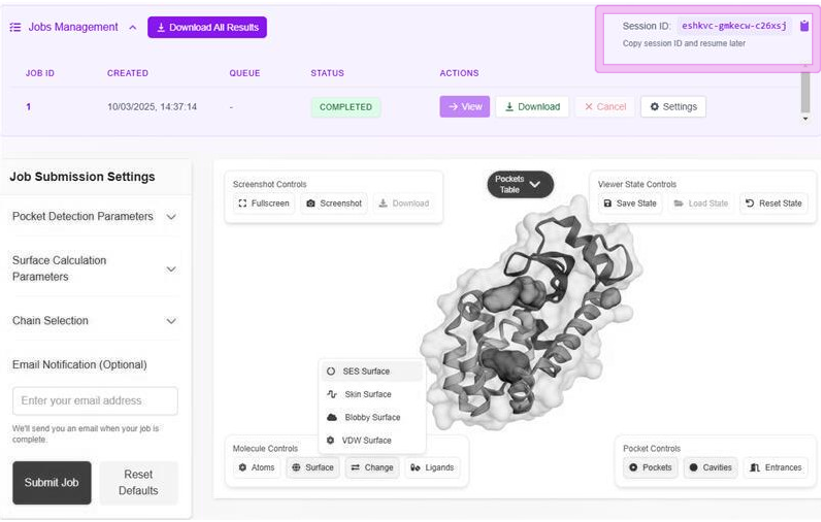
Surface Types
Connolly-Richards
Classical molecular surface representation
Skin Surface
Alternative smooth molecular surface
Blobby Surface
Gaussian-based representation
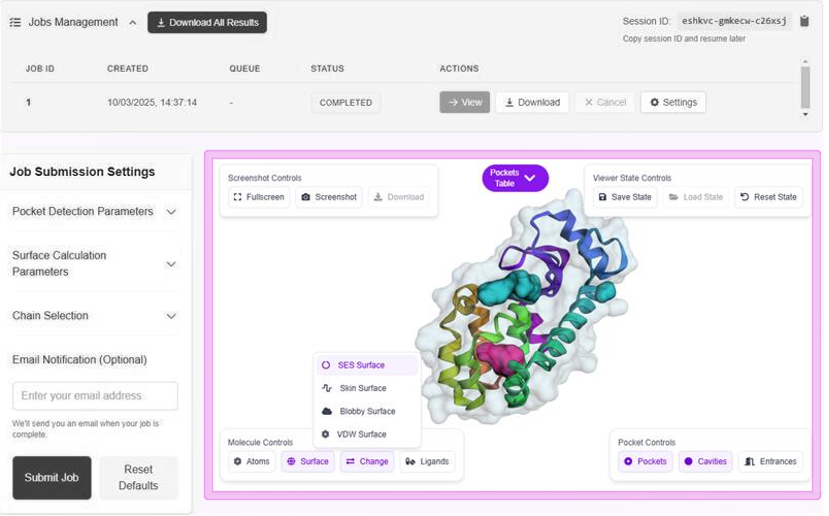
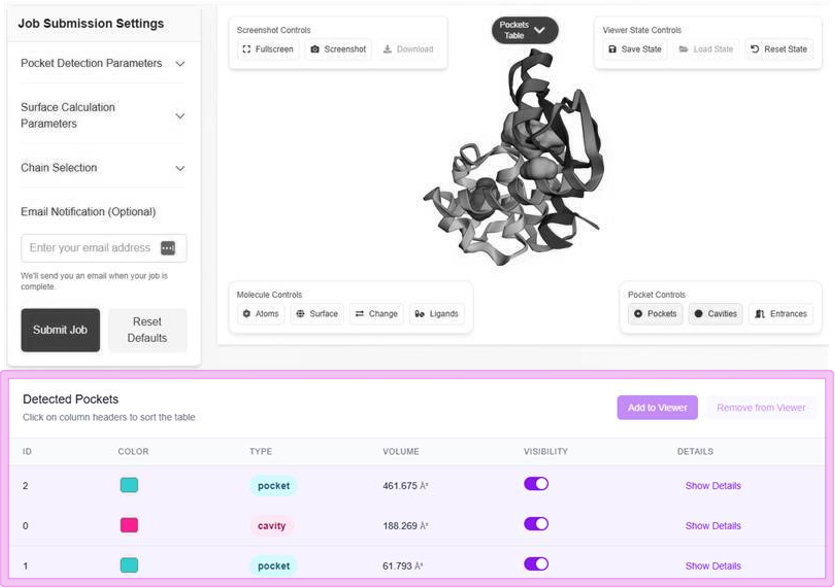
Pocket Detection
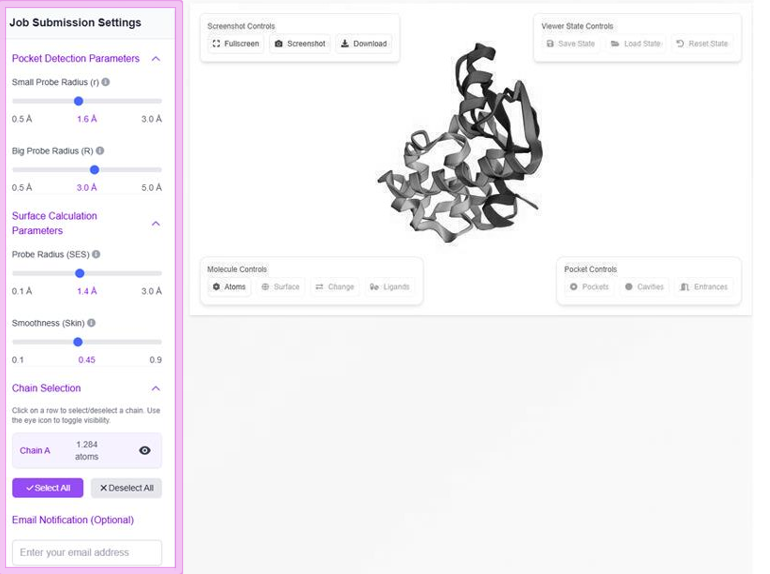
The identification of internal cavities and pockets uses an intuitive volumetric difference approach, using two probe radii to define regions between solvent-excluded surfaces (SESs).
Probe Sizes
- Large probe with radius R (default: 3.0 Å)
- Smaller probe with radius r (default: 1.4 Å)
Effects
- Higher R value identifies shallow pockets
- Higher r value smooths inner surface gaps
Technical Details
Implementation
NanoShaper is written in portable and extensible C++, making it adaptable for various applications. Users can analyze PDB files, import MSMS files, and generate molecular surfaces such as skin, blobby, and SES surfaces.
Surface Area Calculation
The surface area calculation is performed using a parallel Marching Cubes algorithm. When voids are removed, the triangulation reflects this by producing a consistent surface representation without cavities.
Volume Estimation Process
The volume estimation process involves a sophisticated triple ray-casting approach:
- Rays are cast from all the three coordinated planes (xy, xz, yz)
- Each group of rays cast by a plane provides an independent volume estimation
- The three measurements are averaged for maximum accuracy
References
- Decherchi, Sergio, and Walter Rocchia. "A general and Robust Ray-Casting-Based Algorithm for Triangulating Surfaces at the Nanoscale." Plos One 8, no. 4 (2013): 1-15. https://doi.org/10.1371/journal.pone.0059744.
- Krasowski, Agata, Daniel Muthas, Aurijit Sarkar, Stefan Schmitt, and Ruth Brenk. "DrugPred: a structure-based approach to predict protein druggability developed using an extensive nonredundant data set." Journal of chemical information and modeling 51, no. 11 (2011): 2829-2842.
- Nicholas Rego, David Koes, 3Dmol.js: molecular visualization with WebGL, Bioinformatics, Volume 31, Issue 8, April 2015, Pages 1322–1324, https://doi.org/10.1093/bioinformatics/btu829.
Cite NanoShaperWeb
If you use NanoShaperWeb in your research, please cite:
BibTeX Format
author={Abate, Carlo and Serra, Eleonora and Rocchia, Walter and Cavalli, Andrea and Decherchi, Sergio},
title={NanoShaperWeb: Molecular Surface and Pocket Detection Made Visual},
journal={Journal of Chemical Information and Modeling},
year={2025},
month={Jun},
day={30},
publisher={American Chemical Society},
issn={1549-9596},
doi={10.1021/acs.jcim.5c00821},
url={https://doi.org/10.1021/acs.jcim.5c00821}
}
ACM Format
Contact
For questions and support, please contact:
hpc_servicedesk@iit.it